This page was produced as an assignment for Genetics 677
Experiment Proposal
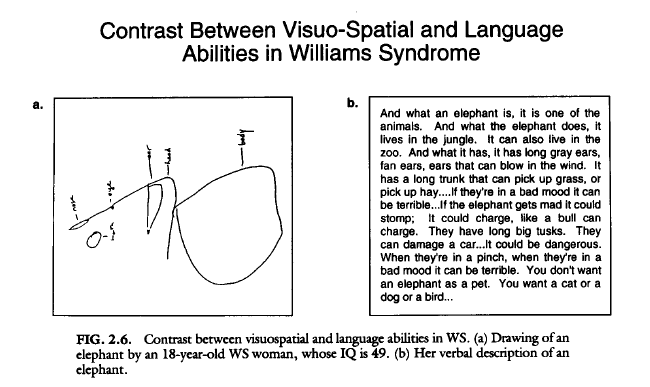
Williams syndrome is a disease characterized by a combination of physical and behavioral symptoms. One of the most interesting cognitive disabilities is their lack of visuospatial cognition, as seen in Fig 1. In this figure (also seen on the homepage), taken from a study done by Belugi et al. in 1994, an 18 year old woman with an IQ of 49 was asked to draw and elephant and then describe it. As you can see in the figure, there is a drastic difference between her writing and drawing ability. The drawing is an example of this lack of visuospatial cognition.
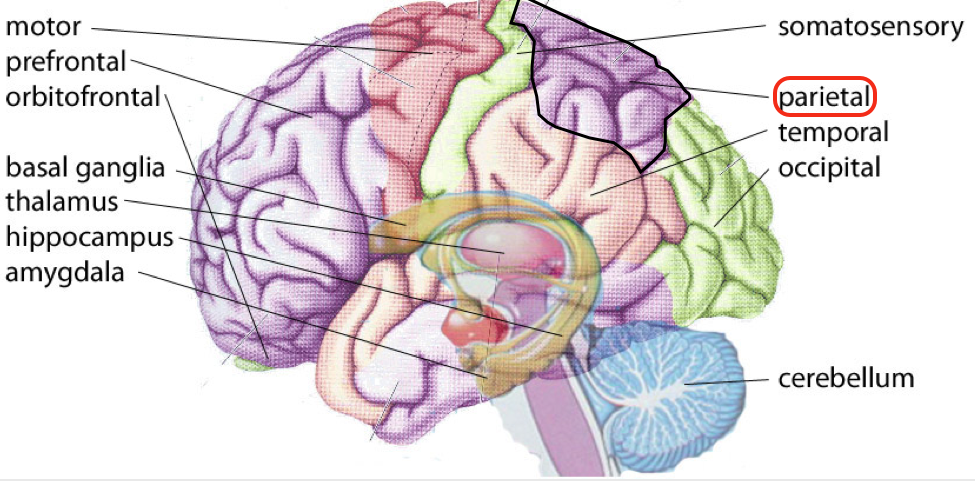
I chose to research LIMK1 out of the ~20 genes deleted from the q arm of chromosome 7 because I am interested in how LIMK1 affects visuospatial cognition. Previous studies have knocked out LIMK1 in mice and have observed this phenotype (1). After doing a little research, I found that visuospatial cognition is linked to the parietal lobe of the brain (Fig 2). This led me to wonder how deletion of LIMK1 results in this phenotype.
When researching, I found that LIMK1 is responsible for actin cytoskeleton organization, among other biological processes related to axon guidance, extension, and neural projection. These are all processes important for brain development, but I will mostly focus on actin cytoskeleton organization. So, you’re probably wondering how the actin cytoskeleton affects nervous system development. The answer is… neuron pathfinding! Nervous system development is dependent on this. Neuron pathfinding is the process by which axons are guided to their target cells by specific extracellular cues. The ability of neurons to move is possible by the growth cone. Motility of the growth cones is dependent on rapid reorganization of the actin cytoskeleton. Previous studies have found that LIMK1 regulates the actin cytoskeleton organization in the hippocampus. Fig 3 shows WT neurons on the left, and LIMK1 knockout (KO) neurons on the right, observed in the study by Meng et al. (1). As you can see, there is less branching out in the LIMK1 KO neuron. Also, there is a lack of growth cones (as shown by the black arrows) and an abnormal distribution of cofilin (stained as greenish yellow). As seen by the irregular distribution of cofilin, it is important not only to consider what happens with LIMK1 is knocked out, but also how this effects expression of the proteins it interacts with.
|
Figure 1. Visuo-spatial ability compared to linguistic ability. From Bellugi, U., 1994.(2) Click for larger image.
|
|
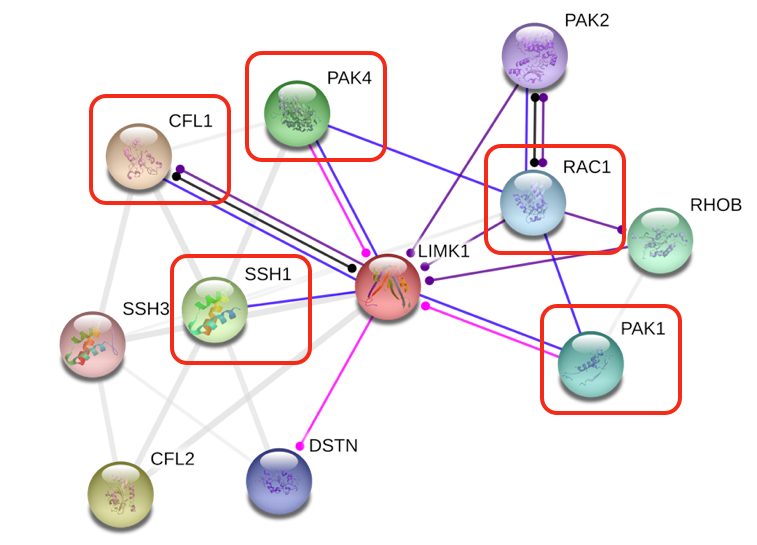
In the red boxes of Fig 4, I have shown which proteins are particularly expressed in the brain: -CLF1 (cofilin): Intracellular actin-modulating protein that binds and depolymerizes F-actin. Important in cell motility and the actin cytoskeleton. -SSH1: Part of the ADF/cofilin family. It is a protein phosphatase that regulates actin filament dynamics and also interacts with cofilin. -PAK1: Serine/threonine p21-activating kinase. Critical effector that links Rho GTP-ases to the cytoskeleton. It is involved in cytoskeleton reorganization and nuclear signaling. Actions of PAK1 are mediated by cofilin-dependent actin regulation. -PAK4: Mediator of filipodia formation. It also plays a role in reorganization of the actin cytoskeleton. PAK4 phosphorylates and inactivates SSH1, which leads to decreased cofilin activity and may lead to stabilization of actin filaments. -Rac1: GTP-ase that belongs to the RAS superfamily and is responsible for control of cell growth, cytoskeletal reorganization, and the activation of protein kinases. |
Figure 3. WT neurons on left and LIMK1 KO neurons on right. Image from Meng Y, Zhang Y, Tregoubov V, Janus C, Cruz L, Jackson M, Lu WY, MacDonald JF, Wang JY, Falls DL, and Jia Z. Abnormal spine morphology and enhanced LTP in LIMK-1 knockout mice. Neuron 35: 121–133, 2002. Click for larger image.
|
|
Figure 4. STRING network of proteins that interact with LIMK1 in humans. Red boxes indicate proteins that are found in relatively high amounts in the brain (STRING).
|
It is also important to consider the binding domain of LIMK1, as well as its interacting proteins (Fig 5 and 6). These binding domains determine how the proteins interact and how they could potentially be important for chemical screens in the future. LIMK1 has (Fig 5): -2 LIM domains: These are zinc binding domains. LIM domains can also bind protein partners via tyrosine-containing motifs. They are found in many key regulators of developmental pathways. -PDZ domain: Found in diverse signaling proteins. It binds C-terminal polypeptides. -Protein tyrosine kinase domain: Functions as on/off switch for many cellular functions and is important for transmembrane signaling. For more information, see (Motifs and Domain) Figure 5. Domains of LIMK1 (PFAM).
|
|
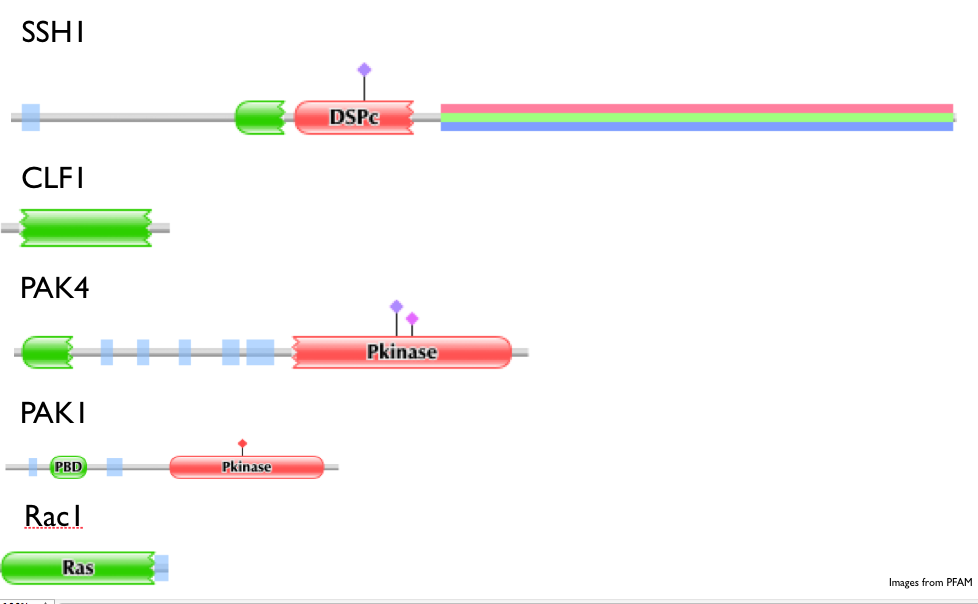
Figure 6. Domains of interacting proteins with LIMK1 (PFAM).
|
Domains of some interacting proteins with LIMK1 (Fig 6): such as: -PAK1 and PAK4: Contain protein kinase domains, which function as on/off switches for cellular processes as well, and act through enzymes that move phosphate groups to proteins. This domain as been shown to function in embryonic development and abnormal phosphorylation can result in disease. -SSH1: Contains a DSPc domain, which is a protein tyrosine phosphatase. This is responsible for removing phosphate groups from phosphorylated tyrosine residues on proteins. It is important for cell growth, differentiation, proliferation, and transformation. -CLF1: Has been shown to interact with SSH1. Contains a ADF-H domain, which promotes rapid filament turnover in cells by depolymerizing actin filaments. -Rac1: Contains a Ras domain, which is involved in cellular signal transduction and can be switched on by incoming signals and then can switch on transcription of other proteins. This ultimately turns genes on related to cell growth, differentiation, and survival. |
As you can see, these proteins and their interactions are important for cell growth and development, and many interact with one another. Several are involved in maintenance and formation of the cytoskeleton, which leads me to hypothesize that deletion of LIMK1 will alter the actin cytoskeleton organization of the parietal neurons. Again, the phenotype of visuospatial cognition has been linked to the parietal lobe, which is why I chose to study these neurons specifically.
I first plan to establish where LIMK1 and its interacting proteins localize in these neurons, and then establish the role of cofilin in actin cytoskeleton organization. I chose cofilin specifically because it interacts with LIMK1 as well as many of the other proteins that interact with LIMK1. To study my question, I will use a mouse as my model organism. As you can see from the phylogenetic tree (see phylogenetic tree link in the toolbar at the top of the page), LIMK1 is very similar in humans and mice, with an E-value of 2x10^-79. Additionally, LIMK1 in humans and mice interacts with many of the same proteins (which can be seen in the presentation file below).
For my first experiment, I plan to create transgenic mice with a LIMK1 deletion (knockout LIMK1). This generates a partial Williams Syndrome phenotype, as shown by a previous study done by Meng et al. Then, in both the wild type and LIMK1 knockout mice, I will use immunofluoresence to determine where LIMK1 localizes, as well as where all of its interacting proteins mentioned previously. This will help determine:
1) Where do the proteins localize?
2) Do they localize to the same areas of the parietal neurons when LIMK1 is deleted?
I will then use a microarray to determine expression of these interacting proteins when LIMK1 is present and when it is knocked out. This will allow me to determine which proteins are affected by the absence of LIMK1, and will suggest a key interaction between LIMK1 and the protein that is missing. One downfall of these techniques is that it is not possible to determine if other proteins are able to function as LIMK1 functions, which may result in normal expression and function of LIMK1’s interacting proteins.
1) Where do the proteins localize?
2) Do they localize to the same areas of the parietal neurons when LIMK1 is deleted?
I will then use a microarray to determine expression of these interacting proteins when LIMK1 is present and when it is knocked out. This will allow me to determine which proteins are affected by the absence of LIMK1, and will suggest a key interaction between LIMK1 and the protein that is missing. One downfall of these techniques is that it is not possible to determine if other proteins are able to function as LIMK1 functions, which may result in normal expression and function of LIMK1’s interacting proteins.
|
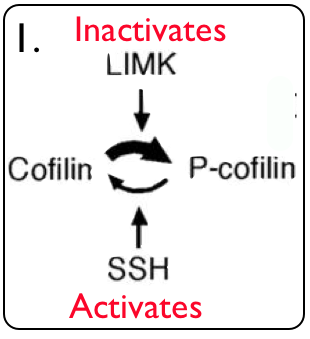
For my second experiment, I plan to further observe how cofilin is affected when LIMK1 is knocked out. LIMK1 phosphorylates cofilin, rendering it inactive (Fig 7). SSH1 activates cofilin through dephosphorylation. I will create transgenic knockout LIMK1 mice, and then determine protein expression using a microarray. I expect that cofilin will be hyperactivated. I will use GFP to determine localization, which I predict will be more in the dendrites of the parietal neurons based on the previous experiment in the hippocampus (name experiment). I will then use immunohistochemistry to determine the amounts of phosphorylated cofilin, which is the amount inactivated. I expect this to be lower in the LIMK1 KO mice. I will not be able to measure post synaptic density with these experiments, but I will hopefully be able to determine if cofilin’s effect on actin filament dynamics results in the change in morphology of dendritic spines or if the deletion of LIMK1 has another unknown effect that results in change in morphology.
|
Figure 7. LIMK1, Cofilin, and SSH1 interaction (4).
|
Presentation
Below is a link to my final presentation in PowerPoint format. It includes the information on this page as well as some extra diagrams, such as the Mus musculus STRING network. One error in the presentation is the use of RNAi to make a knockout LIMK1 mouse. Instead, I plan to use a transgenic mouse as discussed earlier.
| williams_syndrome_-_final_presentation.ppt | |
| File Size: | 3419 kb |
| File Type: | ppt |
References
1. Meng Y, Zhang Y, Tregoubov V, Janus C, Cruz L, Jackson M, Lu WY, MacDonald JF, Wang JY, Falls DL, and Jia Z. Abnormal spine morphology and enhanced LTP in LIMK-1 knockout mice. Neuron 35: 121–133, 2002.
2. Bellugi, U.,Wang, P. P., & Jernigan, T. L. (1994).Williams syndrome: An unusual neuropsychological profile. In S. H. Broman & J. Grafman (Eds.), A typical cognitive deficits in developmental disorders: Implications for brain function (pp. 23–56). Hillsdale, NJ: Lawrence Erlbaum.
3.http://neuralmodel.net/library/brain/parietal_cortex.htm
4. Meng Y, Takahashi H, Meng J, Zhang Y, Lu G, Asrar S, Nakamura T, Jia Z. Regulation of ADF/cofilin phosphorylation and synaptic function by LIM-kinase. Neuropharmacology. 2004;47:746–754.
5.Samiere PD, Bamburg JR. Head, neck, and spines: a role for LIMK-1 in the hippocampus. Neuron. 2002;35:3-5/
All information about domains is from PFAM. All information about interacting proteins is from Uniprot.
The header image was made using images from the following URLS:
http://learn.genetics.utah.edu/content/disorders/whataregd/williams/
http://whoneedswho.areavoices.com/2011/01/31/hello-world/
http://mindbodyshift.wordpress.com/2010/05/18/an-unquenchable-thirst-for-love-the-paradox-of-living-with-williams-syndrome/
http://www.williams-syndrome.org/diagnosing-williams-syndrome/diagnosing-williams-syndrome
http://www.science3point0.com/genegeek/2010/11/08/human-chromosomes-and-karyotype/
http://www.pdb.org/pdb/explore/explore.do?structureId=3S95
1. Meng Y, Zhang Y, Tregoubov V, Janus C, Cruz L, Jackson M, Lu WY, MacDonald JF, Wang JY, Falls DL, and Jia Z. Abnormal spine morphology and enhanced LTP in LIMK-1 knockout mice. Neuron 35: 121–133, 2002.
2. Bellugi, U.,Wang, P. P., & Jernigan, T. L. (1994).Williams syndrome: An unusual neuropsychological profile. In S. H. Broman & J. Grafman (Eds.), A typical cognitive deficits in developmental disorders: Implications for brain function (pp. 23–56). Hillsdale, NJ: Lawrence Erlbaum.
3.http://neuralmodel.net/library/brain/parietal_cortex.htm
4. Meng Y, Takahashi H, Meng J, Zhang Y, Lu G, Asrar S, Nakamura T, Jia Z. Regulation of ADF/cofilin phosphorylation and synaptic function by LIM-kinase. Neuropharmacology. 2004;47:746–754.
5.Samiere PD, Bamburg JR. Head, neck, and spines: a role for LIMK-1 in the hippocampus. Neuron. 2002;35:3-5/
All information about domains is from PFAM. All information about interacting proteins is from Uniprot.
The header image was made using images from the following URLS:
http://learn.genetics.utah.edu/content/disorders/whataregd/williams/
http://whoneedswho.areavoices.com/2011/01/31/hello-world/
http://mindbodyshift.wordpress.com/2010/05/18/an-unquenchable-thirst-for-love-the-paradox-of-living-with-williams-syndrome/
http://www.williams-syndrome.org/diagnosing-williams-syndrome/diagnosing-williams-syndrome
http://www.science3point0.com/genegeek/2010/11/08/human-chromosomes-and-karyotype/
http://www.pdb.org/pdb/explore/explore.do?structureId=3S95
This page was last edited by Natalie DeCheck on May 23, 2012