Protein Interactions
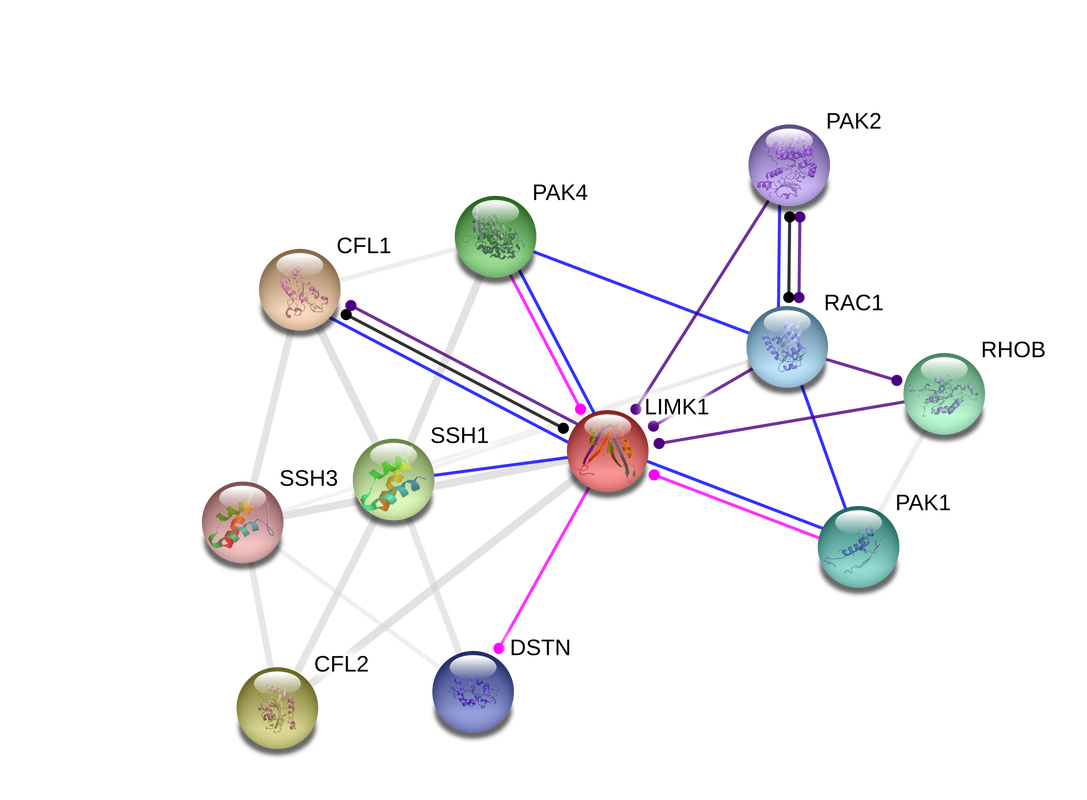
_LIMK1 has the highest levels of expression in both the adult and fetal nervous system, with lower levels of expression in the heart and skeletal muscle (Uniprot). In Fig 1, the STRING network, or protein-protein interactions between LIMK1 and other proteins, is depicted. LIMK1 is a serine/threonine protein kinase and acts downstream of serveral Rho familiy GTPase signal transduction pathways (1).
It is activated by upstream kinases, such as the following, via phosphorylation on a threonine residue (1): -ROCK1 on Thr-508 -PAK1 on Thr-508 -PAK4 (increases LIMK1's ability to phosphorylate cofilin) LIMK1 is also activated by phosphorylation at Ser-323 by MAPKAPK2 during activation of VEGFA-induced signaling. This promotes actin reorganization and cell migration (1). LIMK1 phosphorylates and inactivates the following actin binding/depolymerizing factors (1): -CFL1 -CFL2 -DSTN This results in the stabilization of the actin cytoskeleton, which regulates cell motility, the cell cycle, and cell differentiation. |
Figure 1. STRING network of LIMK1 in humans (2).
|
Post-translational modifications
_- inactivated by SSHI via dephosphorylation (1).
- ubiquination by RNF6 leads to degradation by the 26S proteasome. This modulates LIMK1 levels in the growth cone, regulating axonal outgrowth (1).
- ubiquination by RNF6 leads to degradation by the 26S proteasome. This modulates LIMK1 levels in the growth cone, regulating axonal outgrowth (1).
Analysis
Many of these interactions are important for actin cytoskeleton reorganization and movement in neurons. When LIMK1 is deleted, CFL1, CFL2, and DSTN are not phosphorylated and therefore cannot be inactivated by LIMK1. Little is known about what happens to the expression of these proteins when LIMK1 is deleted, which I propose to examine in my experiment .
References:
1. http://www.uniprot.org/uniprot/P53667
2.http://string-db.org/
The header image was made using images from the following URLS:
http://learn.genetics.utah.edu/content/disorders/whataregd/williams/
http://whoneedswho.areavoices.com/2011/01/31/hello-world/
http://mindbodyshift.wordpress.com/2010/05/18/an-unquenchable-thirst-for-love-the-paradox-of-living-with-williams-syndrome/
http://www.williams-syndrome.org/diagnosing-williams-syndrome/diagnosing-williams-syndrome
http://www.science3point0.com/genegeek/2010/11/08/human-chromosomes-and-karyotype/
http://www.pdb.org/pdb/explore/explore.do?structureId=3S95
1. http://www.uniprot.org/uniprot/P53667
2.http://string-db.org/
The header image was made using images from the following URLS:
http://learn.genetics.utah.edu/content/disorders/whataregd/williams/
http://whoneedswho.areavoices.com/2011/01/31/hello-world/
http://mindbodyshift.wordpress.com/2010/05/18/an-unquenchable-thirst-for-love-the-paradox-of-living-with-williams-syndrome/
http://www.williams-syndrome.org/diagnosing-williams-syndrome/diagnosing-williams-syndrome
http://www.science3point0.com/genegeek/2010/11/08/human-chromosomes-and-karyotype/
http://www.pdb.org/pdb/explore/explore.do?structureId=3S95
Last updated by Natalie Decheck on May 22, 2012